ALZHEIMER'S DISEASE
IAG’s team contributes into the design and execution of clinical trials. Specifically, in Alzheimer’s Disease (AD), we are supporting the considerable amount of effort dedicated to finding new effective therapeutic agents for patients, with several symptomatic and disease-modifying therapies currently being evaluated in clinical trials.
Collaboratively, with leading academics and medical institutions, we also work on developing and validating biomarkers and companion digital diagnostic markers, which may assist in early diagnosis, trial enrichment, and objective measurement of disease progression.
Alzheimer’s disease is a brain disorder that slowly destroys memory and thinking skills, and eventually, the ability to carry out the simplest tasks.
In relation to patient selection for drug development programs, it is important to note that the clinical presentation of AD is heterogeneous and overlaps with other conditions. Brain MRI uses different sequences and contrasts to study brain structure and function while PET imaging use ionising radioactive ligands to quantitatively measure receptors, transporters or enzymes at nanomolecular level with high specificity and power of resolution.
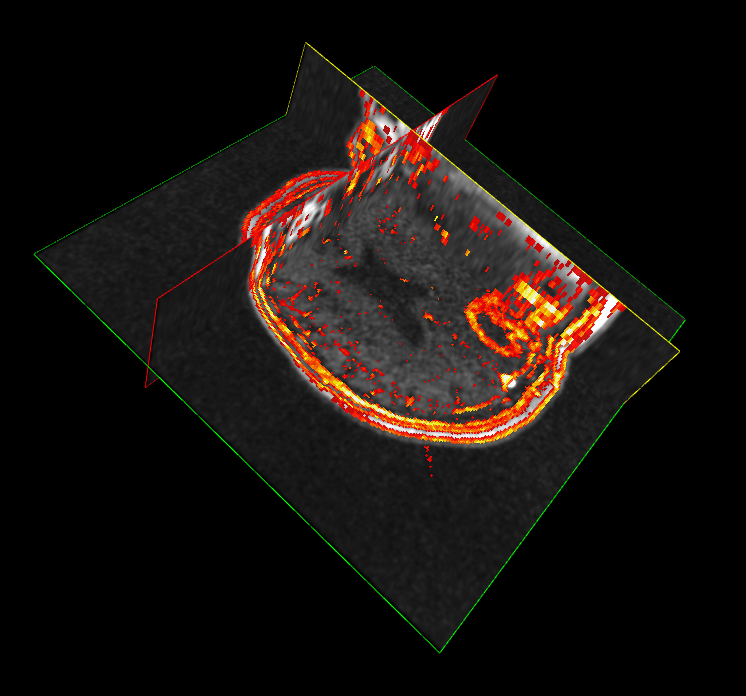
Imaging of the brain in patients with AD can increase the accuracy of differential diagnosis, patient selection and assessment of the disease progression.
- MRI / CT: computed tomography (CT) and then magnetic resonance imaging (MRI) are used diagnostically to rule out other causes of dementia
- Structural MRI is useful to assess volumetric changes of the brain and for measuring the degree and distribution of brain atrophy.
A typical MRI dataset from AD trial would have: FLAIR, T2, T2*, DWI (trace), DWI (ADC), T1 & Post-Gadolinium T1
- Advanced MRI modalities, including diffusion-weighted imaging (DWI), spectroscopy, arterial spin labelling (ASL) and resting-state functional MRI can assist in detection of and discrimination of AD cases from other forms of dementia.
- Functional MRI measures blood oxygenation-level dependent (BOLD) signal, which is sensitive to localised changes in levels of blood oxygenation in brain regions that are activated.
- Imaging with PET or SPECT are powerful methods to detect in vivo changes in the brain at molecular level.
- Structural and functional MRI and positron emission tomography (PET) studies of cerebral metabolism with fluoro-deoxy-d-glucose (FDG) and amyloid tracers have shown characteristic changes in the brains of patients with AD, and in prodromal and even presymptomatic states that can help rule-in the AD pathophysiological process.
- Using PET imaging with 18F-fludeoxyglucose (18F-FDG), patterns of regional brain metabolism can be measured using a map of regional abnormalities of known disease-specific templates obtained in established patient cohorts.
At IAG, we believe that the most important requirements for introducing quantitative MRI endpoints in the design of AD trials are the accuracy & reproducibility of the analysis techniques.
The processes and operational constraints should be scalable for a successful deployment in the context of large multicentre clinical trials. The biggest scientific and operational challenge is in making use of advanced quantitative image processing techniques in a highly regulated environment (FDA’s 21 CFR Part 11 & DYNAMIKA’s ISO13485 software validation guidelines).
Our operational teams are here to ensure that we are maximizing the quality of MRI data, minimizing intra/ inter-site variability, providing vendor & model-specific MRI acquisition parameters.
DYNAMIKA supports the ongoing quality control of MRI data performed within short turnaround times and we consider it to be one of the most crucial tasks to increase the accuracy & reproducibility of image processing techniques.
In our studies, we use American College of Radiology (ACR) phantom for initial site qualification and for monitoring the image quality over time.
The data collection is fully transparent and is done using web based electronic image transfer within DYNAMIKA.
WHAT IS ATLAS-BASED SEGMENTATION:
An atlas-based segmentation isa technique that is used for the detection of brain, ventricles, hippocampus and other brain subcortical structures, using atlases or a collection of brain scans. The quality of brain structures delineation at baseline is of major importance for subsequent image analysis steps.
WHAT IS BRAIN, HIPPOCAMPAL & VENTRICULAR ATROPHY
QUANTIFICATION?
A registration-based (also known as alignment of images) or segmentation based (detection of structures) can be used for the quantification of brain atrophy. Test-retest reproducibility results are very important to assess accuracy of a technique. This variability or “measurement error” inherent to the quantification process is well below the expected mean annual change of total brain volume in healthy elderly controls.
IAG’s team has deep understanding of challenges associated with design and execution of AD trials.
We understand that optimal clinical trial design is crucial. Chosen imaging modality and associated image analysis will help to prove the efficacy of the therapy. We will recommend the optimal imaging and help selecting the trial endpoints. Once the trial is designed, IAG’s team will select and train the sites, assist with imaging data collection and review. The complex nature of the quantitative endpoints usually requires the use of automated image processing capabilities, enabling high-throughput analysis. IAG’s DYNAMIKA platform is well suited to multi-centre trials, which require regulatory compliance and robust execution.
We make sure that any developed designs are supported by practical approaches that are robust enough for variations in image quality (especially in multicentre contexts, where different MRI scanner manufacturers & models are used.
Our team of board-certified Neuroradiologists independently assess both native and processed MRI data for the evaluation of eligibility, safety and efficacy endpoints. The readers are trained and read simultaneously using our platform, which supports automated adjudication. Conduct of remote read sessions can significantly increase the efficiency of central reading activities while minimizing the inherent costs.
The image evaluation results are made available at the sponsor’s site in a real-time manner to facilitate the tracking of operations and allowing fast decision making when it comes to patient monitoring, in case of safety findings.
About IAG, Image Analysis Group
IAG is a unique partner to life sciences companies developing new treatment and driving the hope of the up-coming precision medicine. IAG leverages expertise in medical imaging and the power of DYNAMIKA™, our proprietary cloud-based platform, to de-risk clinical development and deliver lifesaving therapies into the hands of patients much sooner. IAG provides early drug efficacy assessments, smart patient recruitment and predictive analysis of advanced treatment manifestations, thus lowering investment risk and accelerating study outcomes.
Acting as imaging Contract Research Organization, IAG’s experts also recognize the significance of a comprehensive approach to asset development. They actively engage in co-development projects with both private and public sectors, demonstrating a commitment to cultivating collaboration and advancing healthcare solutions.
Contact our expert team: imaging.experts@ia-grp.com
Selected Publications by IAG’s Team:
- Hajiesmaeili, M., Dehmeshki, J., Bagheri Nakhjavanlo, Band Ellis, T. J. (2014) Initialisation of 3D level set for hippocampus segmentation from volumetric brain MR images. In: Sixth International Conference on Digital Image Processing; 5 Apr 2014, Athens, Greece. (SPIE Proceedings, no. 9159)
- Hajiesmaeili, Maryam, Bagherinakhjavanlo, Bashir, Dehmeshki, Jamshidand Ellis, Tim (2012) Segmentation of the hippocampus for detection of Alzheimer’s disease. In: International Symposium on Visual Computing : 8th International Symposium; 16 – 18 Jul 2012, Crete, Greece.
- Hajiesmaeili, M., Dehmeshki, J. and Ellis, T. J. (2014) Analysis of Partial Volume Effects on Accurate Measurement of the Hippocampus Volume. Journal of Electronic Science and Technology, 2, ISSN (print) 1672-6464
- Traboulsee, A., Dehmeshki, J., Peters, K.R., Griffin, C.M., Brex, P.A., Silver, N.C., Ciccarrelli, O., Chard, D.T., Barker, G.J., Thompson, A.J.and Miller, D.H. (2003) Disability in multiple sclerosis is related to normal appearing brain tissue MTR histogram abnormalities. Multiple Sclerosis, 9(6), pp. 566-573. ISSN (print) 1352-4585
- Dehmeshki, J., Chard, D.T., Leary, S.M., Watt, H.C., Silver, N.C., Tofts, P.S., Thompson, A.J.and Miller, D.H. (2003) The normal appearing grey matter in primary progressive multiple sclerosis. A magnetisation transfer imaging study. Journal of Neurology, 250(1), pp. 67-74. ISSN (print) 0340-5354
- Griffin, C.M., Dehmeshki, Jamshid, Chard, D.T., Parker, G.J., Barker, G.J., Thompson, A.J.and Miller, D.H. (2002) T1 histograms of normal-appearing brain tissue are abnormal in early relapsing-remitting Multiple Sclerosis. Multiple Sclerosis, 8(3), 211-216 (6). ISSN (print) 1352-4585
- Dehmeshki, J., Barker, G.J.and Tofts, P.S. (2002) Classification of disease subgroup and correlation with disease severity using magnetic resonance imaging whole-brain histograms: application to magnetization transfer ratios and multiple sclerosis. IEEE Transactions on Medical Imaging, 21(4), pp. 320-331. ISSN (print) 0278-0062
- Parker, G.J.and Dehmeshki, J. (2002) A level set approach to determining brain region connectivity. In: IWISPA 2000. Proceedings of the First International Workshop on Image and Signal Processing and Analysis. in conjunction with 22nd International Conference on Information Technology Interfaces; 14-15 Jun 2000, Pula, Croatia. (Proceedings of the First International Workshop on Image and Signal Processing and Analysis) ISBN 9539676924
- Toffs, Paul Stephen, Steens, Sca, Dehmeshki, Jamshid, Hofman, Paul, Van Waesberghe, Jan Heinand Van Buchem, Mark (2001) Matching MTR histograms for multi-centre studies. In: ISMRM & ESMRMB Joint Annual Meeting 2001; 21 – 27 Apr 2001, Glasgow, U.K..
- Hatfield, Fraser N.and Dehmeshki, Jamshid (1998) Automatic delineation and 3D visualization of the human ventricular system using probabilistic neural networks. In: Electronic Imaging: Processing, Printing, and Publishing in Color; 18 May 1998, Zurich, Switzerland
Experience: Scoring Systems
3D image registration
MRI signal intensity inhomogeneity correction
Correction of geometrical distortion
Atlas-based segmentation
Quantification of brain, hippocampal & ventricular atrophy
Quantification of intracranial cavity volume
Quantification of FLAIR/T2 hyperintense lesions
Diffusion-Weighted Imaging (DWI)
Diffusion-Tensor Imaging (DTI) data analysis
Incidence of Amyloid Imaging-Related Abnormalities (ARIA)
PET
Experience: Imaging
- MRI
- ARIA
- PET
Publications
Since 2007, over 2000 articles were published to cover scientific discoveries, technology break-throughs and special cases. We list here some critically important papers and abstracts.
Testimonials
Combining our technologies and business advisory services with promising life science companies has yielded spectacular results over the past five years. As a trusted partner to many biotech and pharma companies, IAG’s team is proud to share your words and quotes.